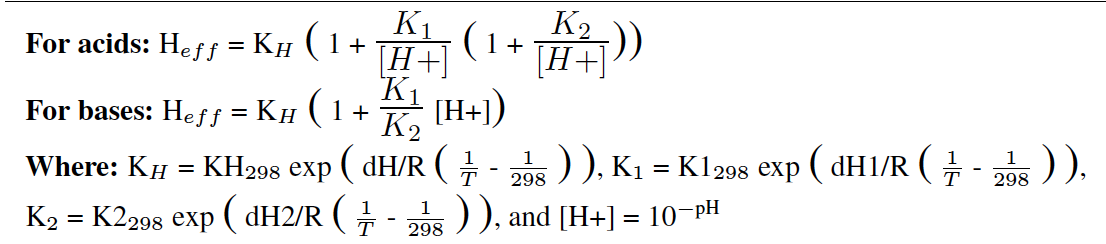The chemical mechanisms and chemistry specific modifications are described in the CAM6 user guide: https://ncar.github.io/CAM/doc/build/html/users_guide/CAM-chem-specifics.html#chemical-mechanisms. A general description of gas-phase chemistry and current developments are provided in the Gas-Phase Chemistry model component description. There are 6 default mechanisms in CESM2 as described in the table below.
| Mechanism | Description | # Species | # Reactions | Mechanism Name | Pre-processor Code |
|---|---|---|---|---|---|
| TSMLT1 | Troposphere, stratosphere, mesosphere, and lower thermosphere | 231 | 583 (433 kinetic, 150 photolysis) | MZ197_TSMLT1_20180423 | pp_waccm_tsmlt_mam4 |
| TS1 | Troposphere and stratosphere | 221 | 528 (405 kinetic, 123 photolysis) | MZ198_TS1-simpleVBS_20180423 | pp_trop_strat_mam4_vbs |
| MA | Middle atmosphere: stratosphere, mesosphere, and lower thermosphere | 98 | 298 (207 kinetic, 91 photolysis) | pp_waccm_ma_mam4 | |
| MAD | Middle atmosphere plus D-region ion chemistry | 135 | 593 (489 kinetic, 104 photolysis) | pp_waccm_mad_mam4 | |
| SC | Specified chemistry for WACCM | 29 | 12 (11 kinetic, 1 photolysis) | pp_waccm_sc_mam4 | |
| CAM | Simplified chemistry for CAM to to allow tropospheric aerosol formation | 32 | 7 (6 kinetic), 1 photolysis) | modal_aero |
The input files and fortran routines for these default mechanisms are located in $CESMROOT/components/cam/src/chemistry/ in the folder specified by the pre-processor code. CAM-Chem and WACCM compsets are defined in section 4.4 and 4.5 of the CAM6 user guide https://ncar.github.io/CAM/doc/build/html/users_guide/atmospheric-configurations.html. All CAM-Chem compsets default to the TS1 chemical mechanism and WACCM compsets default to TSMLT1 chemical mechanism. The TS1 and TSMLT1 chemical mechanisms are described in Emmons et al. (JAMES, 2020).
To review the chemistry used in a specific compset, build your case as described in Run CAM-Chem on Cheyenne. In the $CASEROOT/CaseDocs folder there are 2 files that describe the chemistry chem_mech.in and chem_mech.doc. Both have similar information describing the species and reaction rates. The chem_mech.doc file is in a more human readable format and includes the differential equations defining the production and loss terms of each species. The chem_mech.in file is the version read by the code, and the version you will alter to make changes to the chemical mechanism.
To update the chemistry, modify the chemical mechanism input file by:
- Build your case as described in Run CAM-Chem on Cheyenne.
- Copy the input file (chem_mech.in) from $CASEROOT/CaseDocs folder to another folder and rename the mechanism file (e.g., /home/mech/my_mech.in)
- Then alter this file as needed following these instructions: Altering the Chemical Mechanism Input File
- Then in env_build.xml file, add the following to the CAM_CONFIG_OPTS value "-usr_mech_infile /home/mech/my_mech.in". Note: substitute your own path and mechanism name here.
- You will need to rebuild your case for changes to take effect (see Run CAM-Chem on Cheyenne).
Add emissions for a new species
If a new species is directly emitted, add emissions by:
- Create new emissions files for this species.
- Build your case, copy the ext_frc_specifier and srf_emis_specifier from $CASEROOT/CaseDocs/atm_in to your user_nl_cam file.
- Add your new species and emissions file path to this list.
- ext_frc_specifier are for emissions vertical emissions
- srf_emis_specifier are for surface emissions
- Run ./preview_namelists and confirm the updates appear in your new $CASEROOT/CaseDocs/atm_in file.
Add lower boundary conditions for a new species
For long lived species (e.g., CH4), you can specify the lower boundary conditions instead of having direct emissions from the surface.
- Copy the flbc_file and flbc_list from $CASEROOT/CaseDocs/atm_in to your user_nl_cam file.
- Create a new version of the flbc file that includes your new species.
- Add your new species to the flbc_list.
- Run ./preview_namelists and confirm the updates appear in your new $CASEROOT/CaseDocs/atm_in file.
Include wet and dry deposition of a new species
As of tag cam6_3_071, wet and dry deposition calculations use a netcdf file of Henry's Law data (instead of the data statement described below).
To include deposition of a new species, add the appropriate information (species name, molecular weight, Effective Henry's Law coefficients and dfoxd factor) to the netcdf file, and add the species name to the "drydep_list" and "gas_wetdep_list" (copy existing lists from atm_in and drv_flds_in to user_nl_cam, and edit).
The current "dep_data_file" is listed in atm_in. One way to update it is to use ncdump and ncgen:
- run: ncdump dep_data_orig.nc > dep_data_new.cdl
- edit the cdl text file
- run: ncgen -5 -o dep_data_new.nc dep_data_new.cdl
To add aerosol uptake for a new species
In the TS1 mechanism, organic nitrate aerosol uptake has been included using aerosol uptake coefficients (γ) similar to that used by Fisher et al. 2016.
If you desire to update these aerosol uptake coefficients, follow these instructions for: Adding Aerosol Uptake for Gas-Phase Species.
Note: The code as it is currently written removes the organic nitrates from the gas-phase, but does not have these organic nitrates contribute to secondary organic aerosol formation. Secondary organic aerosol formation is produced through a separate process.
Note: Currently, aerosol uptake of gas-phase compounds only occurs on the following types of aerosol: sulfate, ammonium nitrate, oc2, and secondary organic aerosol.
Wet and dry deposition for CESM2 and earlier:
To wet deposit a new species
There are two methods to wet deposit a new species. Both involve source code changes. After making the changes below, you must rebuild your case for changes to take effect. Wet deposition in CAM-Chem is described in the Wet Deposition model component description.
The first and easiest method is to map the new species to deposit with the same rate as a species already undergoing wet deposition.
- Copy mo_neu_wetdep.F90 from $CESMROOT/components/cam/src/chemistry/mozart/ to your $CASEROOT/SourceMods/src.cam directory.
- Edit the fortran code in mo_neu_wetdep.F90. The example below maps tracer SO2t to SO2.
case( 'SO2t' )
test_name = 'SO2'
- Add species into gas_wetdep_list in user_nl_cam. Note: if the default gas_wetdep_list is not already in your user_nl_cam file, you will need to build your case, copy the default gas_wetdep_list from $CASEROOT/CaseDocs/atm_in, and then add your new species to the end of this list.
The second method is to add a new species to the henry's law table list.
- Copy seq_drydep_mod.F90 from $CESMROOT/cime/src/drivers/mct/shr/ to your $CASEROOT/SourceMods/src.share directory. Note: for older versions of the code, seq_drydep_mod.F90 is in slightly different locations.
- Edit the fortran code in seq_drydep_mod.F90.
- In this code, there are several arrays containing: 1) species names, 2) reactivity factors (f0), 3) henry's law constants, and 4) molecular weights. Add your new species characteristics at the end of each of these arrays. The Henry's law constant table consists of the following 6 columns, respectively: KH298 (M atm-1), dH/R (K), K1298, dH1/R (K), K2298, dH2/R (K). These values are used to calculate the Henry's law constant (KH) and for acids and bases the effective Henry's law constant (Heff) using the following formulas:
Update the variable "maxspc" to be equal or greater than the total number of species you are dry depositing (i.e., the maximum number of species you will dry deposit).
- Update the variable "n_species_table" to the total number of species listed in these arrays.
- In this code, there are several arrays containing: 1) species names, 2) reactivity factors (f0), 3) henry's law constants, and 4) molecular weights. Add your new species characteristics at the end of each of these arrays. The Henry's law constant table consists of the following 6 columns, respectively: KH298 (M atm-1), dH/R (K), K1298, dH1/R (K), K2298, dH2/R (K). These values are used to calculate the Henry's law constant (KH) and for acids and bases the effective Henry's law constant (Heff) using the following formulas:
- Add new species into gas_wetdep_list in user_nl_cam. Note: if the default gas_wetdep_list is not already in your user_nl_cam file, you will need to build your case, copy the default gas_wetdep_list from $CASEROOT/CaseDocs/atm_in, and then add your new species to the end of this list.
To dry deposit a new species
There are also two methods to dry deposit a new species. Both involve source code changes. After making the changes below, you must rebuild your case for changes to take effect. Dry deposition in CAM-Chem is described in the Dry Deposition model component description.
The first and easiest method is to map the new species to deposit with the same rate as a species already undergoing dry deposition.
- Copy seq_drydep_mod.F90 from $CESMROOT/cime/src/drivers/mct/shr/ to your $CASEROOT/SourceMods/src.share directory
- Edit the fortran code in seq_drydep_mod.F90. The example below maps tracer XHNO3 to HNO3. This only works if the "test" species is also simulated.
case( 'XHNO3' )
test_name = 'HNO3'
- Add new species into drydep_list in user_nl_cam. Note: if the default drydep_list is not already in your user_nl_cam file, you will need to build your case, copy the default drydep_list from $CASEROOT/CaseDocs/drv_flds_in, and then add your new species to the end of this list.
The second method is to add a new species to the henry's law table list.
- Copy seq_drydep_mod.F90 from $CESMROOT/cime/src/drivers/mct/shr/ to your $CASEROOT/SourceMods/src.share directory.
- Edit the fortran code in seq_drydep_mod.F90.
- In this code, there are several arrays containing: 1) species names, 2) reactivity factors (f0), 3) henry's law constants, and 4) molecular weights. Add your new species characteristics at the end of each of these arrays. (See updating Wet deposition for a larger description of how these henry's law constants are calculated.
Update the variable "maxspc" to be equal or greater than the total number of species you are dry depositing (i.e., the maximum number of species you will dry deposit).
- Update the variable "n_species_table" to the total number of species listed in these arrays.
- Add new species into drydep_list in user_nl_cam. Note: if the default drydep_list is not already in your user_nl_cam file, you will need to build your case, copy the default drydep_list from $CASEROOT/CaseDocs/drv_flds_in, and then add your new species to the end of this list.
Master lists of species having deposition
The default lists of species with wet or dry deposition are maintained in: $CESMROOT/components/cam/bld/namelist_files/. If a species in these lists is included in the chemical mechanism, then it is included in the default gas_wetdep_list (in atm_in) or drydep_list (in drv_flds_in). Separate lists are kept for aerosol wet and dry deposition (aer_drydep_list and aer_wetdep_list in atm_in).